Planar 3D Transfer Learning for End to End Unimodal MRI Unbalanced Data Segmentation
Martin Kolarik,
Radim Burget,
Carlos M. Travieso-Gonzalez,
Jan Kocica

Auto-TLDR; Planar 3D Res-U-Net Network for Unbalanced 3D Image Segmentation using Fluid Attenuation Inversion Recover
Similar papers
Automatic Semantic Segmentation of Structural Elements related to the Spinal Cord in the Lumbar Region by Using Convolutional Neural Networks
Jhon Jairo Sáenz Gamboa, Maria De La Iglesia-Vaya, Jon Ander Gómez

Auto-TLDR; Semantic Segmentation of Lumbar Spine Using Convolutional Neural Networks
Abstract Slides Poster Similar
A Benchmark Dataset for Segmenting Liver, Vasculature and Lesions from Large-Scale Computed Tomography Data
Bo Wang, Zhengqing Xu, Wei Xu, Qingsen Yan, Liang Zhang, Zheng You

Auto-TLDR; The Biggest Treatment-Oriented Liver Cancer Dataset for Segmentation
Abstract Slides Poster Similar
Deep Recurrent-Convolutional Model for AutomatedSegmentation of Craniomaxillofacial CT Scans
Francesca Murabito, Simone Palazzo, Federica Salanitri Proietto, Francesco Rundo, Ulas Bagci, Daniela Giordano, Rosalia Leonardi, Concetto Spampinato

Auto-TLDR; Automated Segmentation of Anatomical Structures in Craniomaxillofacial CT Scans using Fully Convolutional Deep Networks
Abstract Slides Poster Similar
A Multi-Task Contextual Atrous Residual Network for Brain Tumor Detection & Segmentation
Ngan Le, Kashu Yamazaki, Quach Kha Gia, Thanh-Dat Truong, Marios Savvides

Auto-TLDR; Contextual Brain Tumor Segmentation Using 3D atrous Residual Networks and Cascaded Structures
A Deep Learning Approach for the Segmentation of Myocardial Diseases
Khawala Brahim, Abdull Qayyum, Alain Lalande, Arnaud Boucher, Anis Sakly, Fabrice Meriaudeau

Auto-TLDR; Segmentation of Myocardium Infarction Using Late GADEMRI and SegU-Net
Abstract Slides Poster Similar
FOANet: A Focus of Attention Network with Application to Myocardium Segmentation
Zhou Zhao, Elodie Puybareau, Nicolas Boutry, Thierry Geraud

Auto-TLDR; FOANet: A Hybrid Loss Function for Myocardium Segmentation of Cardiac Magnetic Resonance Images
Abstract Slides Poster Similar
A Systematic Investigation on Deep Architectures for Automatic Skin Lesions Classification
Pierluigi Carcagni, Marco Leo, Andrea Cuna, Giuseppe Celeste, Cosimo Distante

Auto-TLDR; RegNet: Deep Investigation of Convolutional Neural Networks for Automatic Classification of Skin Lesions
Abstract Slides Poster Similar
Do Not Treat Boundaries and Regions Differently: An Example on Heart Left Atrial Segmentation
Zhou Zhao, Elodie Puybareau, Nicolas Boutry, Thierry Geraud

Auto-TLDR; Attention Full Convolutional Network for Atrial Segmentation using ResNet-101 Architecture
Segmentation of Intracranial Aneurysm Remnant in MRA Using Dual-Attention Atrous Net
Subhashis Banerjee, Ashis Kumar Dhara, Johan Wikström, Robin Strand

Auto-TLDR; Dual-Attention Atrous Net for Segmentation of Intracranial Aneurysm Remnant from MRA Images
Abstract Slides Poster Similar
Bridging the Gap between Natural and Medical Images through Deep Colorization
Lia Morra, Luca Piano, Fabrizio Lamberti, Tatiana Tommasi

Auto-TLDR; Transfer Learning for Diagnosis on X-ray Images Using Color Adaptation
Abstract Slides Poster Similar
Transfer Learning through Weighted Loss Function and Group Normalization for Vessel Segmentation from Retinal Images
Abdullah Sarhan, Jon Rokne, Reda Alhajj, Andrew Crichton

Auto-TLDR; Deep Learning for Segmentation of Blood Vessels in Retinal Images
Abstract Slides Poster Similar
Segmenting Kidney on Multiple Phase CT Images Using ULBNet
Yanling Chi, Yuyu Xu, Gang Feng, Jiawei Mao, Sihua Wu, Guibin Xu, Weimin Huang
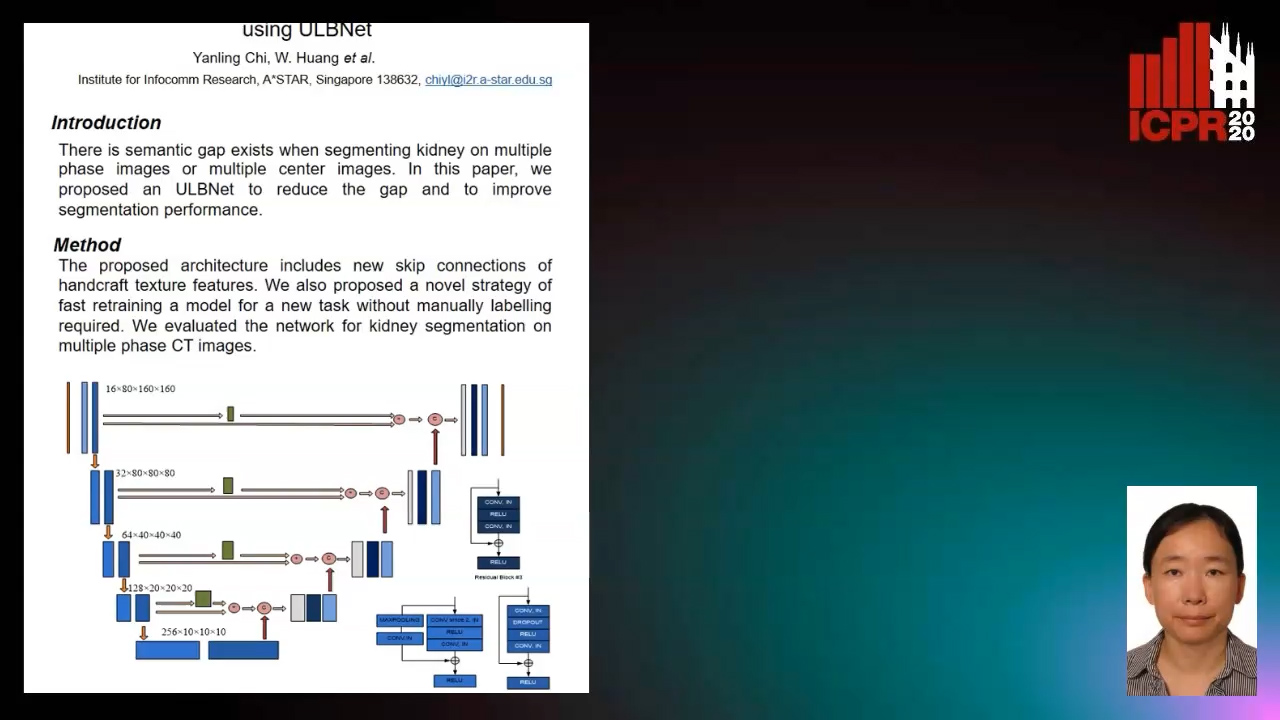
Auto-TLDR; A ULBNet network for kidney segmentation on multiple phase CT images
A Transformer-Based Network for Anisotropic 3D Medical Image Segmentation
Guo Danfeng, Demetri Terzopoulos

Auto-TLDR; A transformer-based model to tackle the anisotropy problem in 3D medical image analysis
Abstract Slides Poster Similar
Learn to Segment Retinal Lesions and Beyond
Qijie Wei, Xirong Li, Weihong Yu, Xiao Zhang, Yongpeng Zhang, Bojie Hu, Bin Mo, Di Gong, Ning Chen, Dayong Ding, Youxin Chen

Auto-TLDR; Multi-task Lesion Segmentation and Disease Classification for Diabetic Retinopathy Grading
3D Medical Multi-Modal Segmentation Network Guided by Multi-Source Correlation Constraint
Tongxue Zhou, Stéphane Canu, Pierre Vera, Su Ruan

Auto-TLDR; Multi-modality Segmentation with Correlation Constrained Network
Abstract Slides Poster Similar
Deep Learning-Based Type Identification of Volumetric MRI Sequences
Jean Pablo De Mello, Thiago Paixão, Rodrigo Berriel, Mauricio Reyes, Alberto F. De Souza, Claudine Badue, Thiago Oliveira-Santos

Auto-TLDR; Deep Learning for Brain MRI Sequences Identification Using Convolutional Neural Network
Abstract Slides Poster Similar
CAggNet: Crossing Aggregation Network for Medical Image Segmentation

Auto-TLDR; Crossing Aggregation Network for Medical Image Segmentation
Abstract Slides Poster Similar
MTGAN: Mask and Texture-Driven Generative Adversarial Network for Lung Nodule Segmentation
Wei Chen, Qiuli Wang, Kun Wang, Dan Yang, Xiaohong Zhang, Chen Liu, Yucong Li
Auto-TLDR; Mask and Texture-driven Generative Adversarial Network for Lung Nodule Segmentation
Abstract Slides Poster Similar
DARN: Deep Attentive Refinement Network for Liver Tumor Segmentation from 3D CT Volume
Yao Zhang, Jiang Tian, Cheng Zhong, Yang Zhang, Zhongchao Shi, Zhiqiang He

Auto-TLDR; Deep Attentive Refinement Network for Liver Tumor Segmentation from 3D Computed Tomography Using Multi-Level Features
Abstract Slides Poster Similar
BCAU-Net: A Novel Architecture with Binary Channel Attention Module for MRI Brain Segmentation
Yongpei Zhu, Zicong Zhou, Guojun Liao, Kehong Yuan

Auto-TLDR; BCAU-Net: Binary Channel Attention U-Net for MRI brain segmentation
Abstract Slides Poster Similar
The DeepHealth Toolkit: A Unified Framework to Boost Biomedical Applications
Michele Cancilla, Laura Canalini, Federico Bolelli, Stefano Allegretti, Salvador Carrión, Roberto Paredes, Jon Ander Gómez, Simone Leo, Marco Enrico Piras, Luca Pireddu, Asaf Badouh, Santiago Marco-Sola, Lluc Alvarez, Miquel Moreto, Costantino Grana

Auto-TLDR; DeepHealth Toolkit: An Open Source Deep Learning Toolkit for Cloud Computing and HPC
Abstract Slides Poster Similar
Confidence Calibration for Deep Renal Biopsy Immunofluorescence Image Classification
Federico Pollastri, Juan Maroñas, Federico Bolelli, Giulia Ligabue, Roberto Paredes, Riccardo Magistroni, Costantino Grana

Auto-TLDR; A Probabilistic Convolutional Neural Network for Immunofluorescence Classification in Renal Biopsy
Abstract Slides Poster Similar
DE-Net: Dilated Encoder Network for Automated Tongue Segmentation
Hui Tang, Bin Wang, Jun Zhou, Yongsheng Gao

Auto-TLDR; Automated Tongue Image Segmentation using De-Net
Abstract Slides Poster Similar
Unsupervised Detection of Pulmonary Opacities for Computer-Aided Diagnosis of COVID-19 on CT Images
Rui Xu, Xiao Cao, Yufeng Wang, Yen-Wei Chen, Xinchen Ye, Lin Lin, Wenchao Zhu, Chao Chen, Fangyi Xu, Yong Zhou, Hongjie Hu, Shoji Kido, Noriyuki Tomiyama

Auto-TLDR; A computer-aided diagnosis of COVID-19 from CT images using unsupervised pulmonary opacity detection
Abstract Slides Poster Similar
Triplet-Path Dilated Network for Detection and Segmentation of General Pathological Images
Jiaqi Luo, Zhicheng Zhao, Fei Su, Limei Guo

Auto-TLDR; Triplet-path Network for One-Stage Object Detection and Segmentation in Pathological Images
Supporting Skin Lesion Diagnosis with Content-Based Image Retrieval
Stefano Allegretti, Federico Bolelli, Federico Pollastri, Sabrina Longhitano, Giovanni Pellacani, Costantino Grana

Auto-TLDR; Skin Images Retrieval Using Convolutional Neural Networks for Skin Lesion Classification and Segmentation
Abstract Slides Poster Similar
Fine-Tuning Convolutional Neural Networks: A Comprehensive Guide and Benchmark Analysis for Glaucoma Screening
Amed Mvoulana, Rostom Kachouri, Mohamed Akil

Auto-TLDR; Fine-tuning Convolutional Neural Networks for Glaucoma Screening
Abstract Slides Poster Similar
Deep Multi-Stage Model for Automated Landmarking of Craniomaxillofacial CT Scans
Simone Palazzo, Giovanni Bellitto, Luca Prezzavento, Francesco Rundo, Ulas Bagci, Daniela Giordano, Rosalia Leonardi, Concetto Spampinato

Auto-TLDR; Automated Landmarking of Craniomaxillofacial CT Images Using Deep Multi-Stage Architecture
End-To-End Multi-Task Learning for Lung Nodule Segmentation and Diagnosis
Wei Chen, Qiuli Wang, Dan Yang, Xiaohong Zhang, Chen Liu, Yucong Li
Auto-TLDR; A novel multi-task framework for lung nodule diagnosis based on deep learning and medical features
A Comparison of Neural Network Approaches for Melanoma Classification
Maria Frasca, Michele Nappi, Michele Risi, Genoveffa Tortora, Alessia Auriemma Citarella

Auto-TLDR; Classification of Melanoma Using Deep Neural Network Methodologies
Abstract Slides Poster Similar
DA-RefineNet: Dual-Inputs Attention RefineNet for Whole Slide Image Segmentation
Ziqiang Li, Rentuo Tao, Qianrun Wu, Bin Li

Auto-TLDR; DA-RefineNet: A dual-inputs attention network for whole slide image segmentation
Abstract Slides Poster Similar
Trainable Spectrally Initializable Matrix Transformations in Convolutional Neural Networks
Michele Alberti, Angela Botros, Schuetz Narayan, Rolf Ingold, Marcus Liwicki, Mathias Seuret

Auto-TLDR; Trainable and Spectrally Initializable Matrix Transformations for Neural Networks
Abstract Slides Poster Similar
Dual Encoder Fusion U-Net (DEFU-Net) for Cross-manufacturer Chest X-Ray Segmentation
Zhang Lipei, Aozhi Liu, Jing Xiao
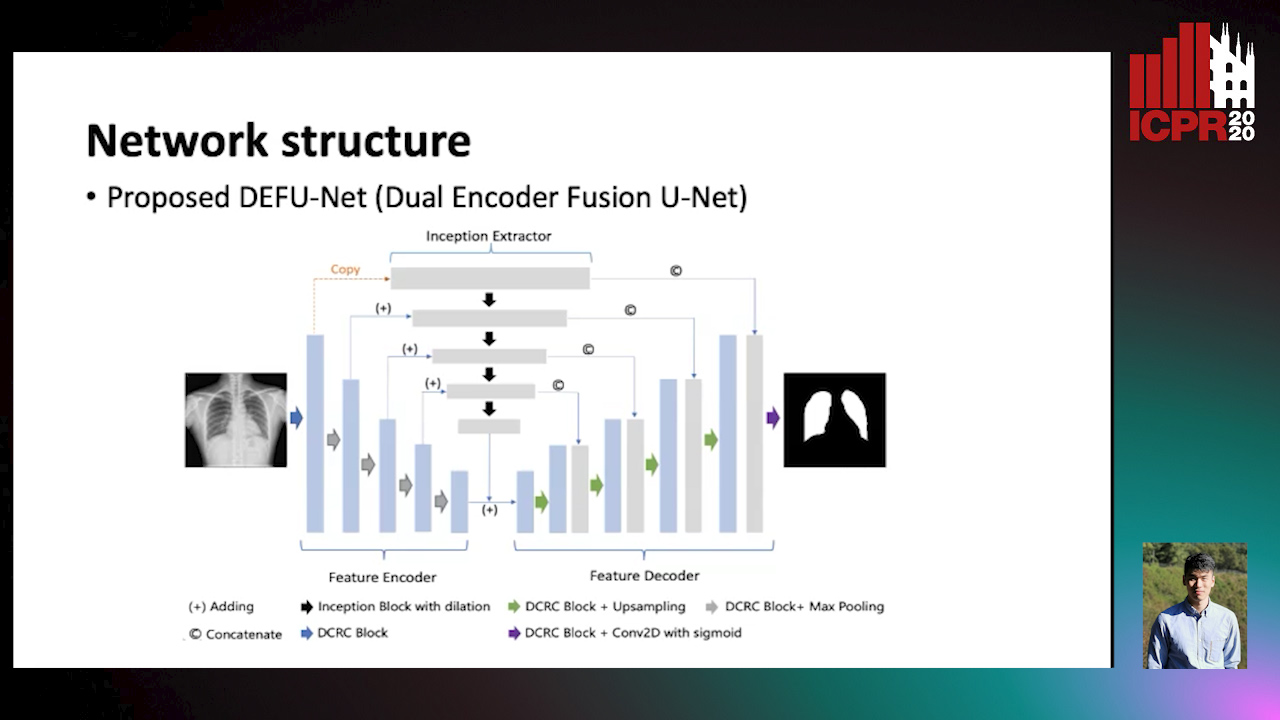
Auto-TLDR; Inception Convolutional Neural Network with Dilation for Chest X-Ray Segmentation
Enhancing Semantic Segmentation of Aerial Images with Inhibitory Neurons
Ihsan Ullah, Sean Reilly, Michael Madden

Auto-TLDR; Lateral Inhibition in Deep Neural Networks for Object Recognition and Semantic Segmentation
Abstract Slides Poster Similar
RescueNet: Joint Building Segmentation and Damage Assessment from Satellite Imagery

Auto-TLDR; RescueNet: End-to-End Building Segmentation and Damage Classification for Humanitarian Aid and Disaster Response
Abstract Slides Poster Similar
Investigating and Exploiting Image Resolution for Transfer Learning-Based Skin Lesion Classification
Amirreza Mahbod, Gerald Schaefer, Chunliang Wang, Rupert Ecker, Georg Dorffner, Isabella Ellinger

Auto-TLDR; Fine-tuned Neural Networks for Skin Lesion Classification Using Dermoscopic Images
Abstract Slides Poster Similar
Leveraging Unlabeled Data for Glioma Molecular Subtype and Survival Prediction
Nicholas Nuechterlein, Beibin Li, Mehmet Saygin Seyfioglu, Sachin Mehta, Patrick Cimino, Linda Shapiro

Auto-TLDR; Multimodal Brain Tumor Segmentation Using Unlabeled MR Data and Genomic Data for Cancer Prediction
Abstract Slides Poster Similar
OCT Image Segmentation Using NeuralArchitecture Search and SRGAN
Saba Heidari, Omid Dehzangi, Nasser M. Nasarabadi, Ali Rezai
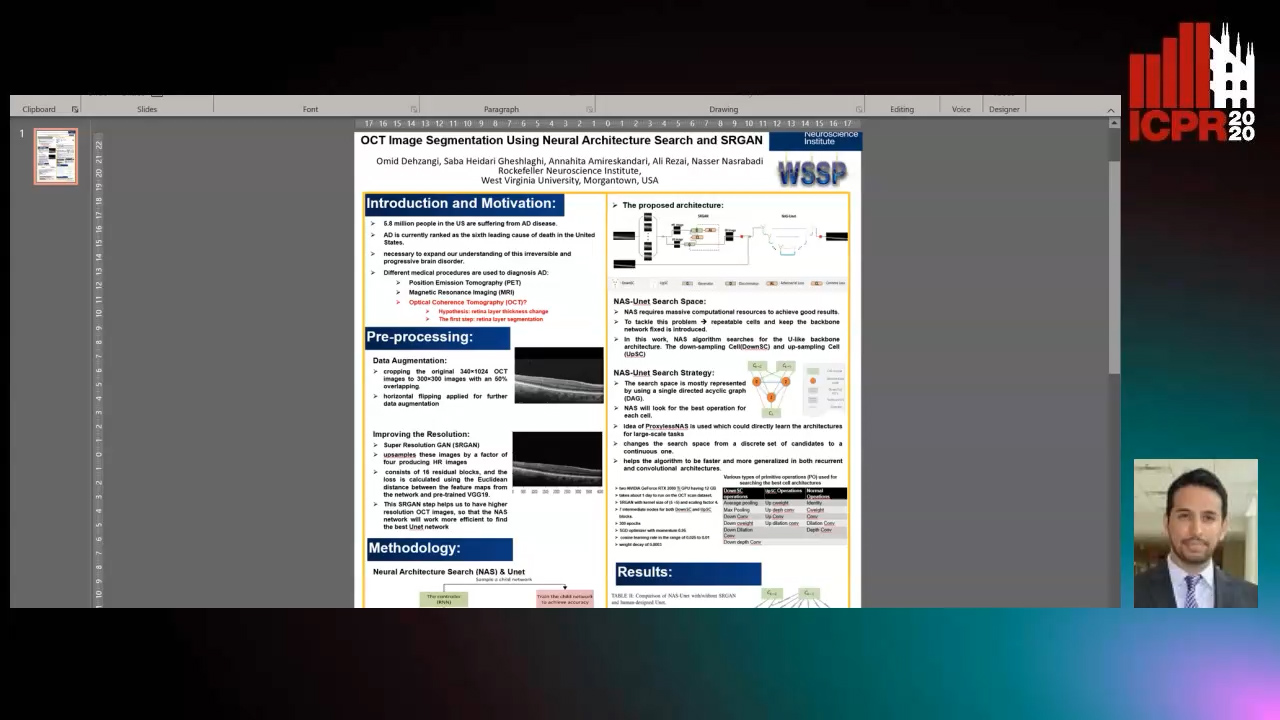
Auto-TLDR; Automatic Segmentation of Retinal Layers in Optical Coherence Tomography using Neural Architecture Search
Fast and Accurate Real-Time Semantic Segmentation with Dilated Asymmetric Convolutions
Leonel Rosas-Arias, Gibran Benitez-Garcia, Jose Portillo-Portillo, Gabriel Sanchez-Perez, Keiji Yanai

Auto-TLDR; FASSD-Net: Dilated Asymmetric Pyramidal Fusion for Real-Time Semantic Segmentation
Abstract Slides Poster Similar
Offset Curves Loss for Imbalanced Problem in Medical Segmentation
Ngan Le, Duc Toan Bui, Khoa Luu, Marios Savvides

Auto-TLDR; Offset Curves Loss for Medical Image Segmentation
EM-Net: Deep Learning for Electron Microscopy Image Segmentation
Afshin Khadangi, Thomas Boudier, Vijay Rajagopal

Auto-TLDR; EM-net: Deep Convolutional Neural Network for Electron Microscopy Image Segmentation
A Lumen Segmentation Method in Ureteroscopy Images Based on a Deep Residual U-Net Architecture
Jorge Lazo, Marzullo Aldo, Sara Moccia, Michele Catellani, Benoit Rosa, Elena De Momi, Michel De Mathelin, Francesco Calimeri

Auto-TLDR; A Deep Neural Network for Ureteroscopy with Residual Units
Abstract Slides Poster Similar
Neural Machine Registration for Motion Correction in Breast DCE-MRI
Federica Aprea, Stefano Marrone, Carlo Sansone

Auto-TLDR; A Neural Registration Network for Dynamic Contrast Enhanced-Magnetic Resonance Imaging
Abstract Slides Poster Similar
Dual Stream Network with Selective Optimization for Skin Disease Recognition in Consumer Grade Images
Krishnam Gupta, Jaiprasad Rampure, Monu Krishnan, Ajit Narayanan, Nikhil Narayan

Auto-TLDR; A Deep Network Architecture for Skin Disease Localisation and Classification on Consumer Grade Images
Abstract Slides Poster Similar
A Heuristic-Based Decision Tree for Connected Components Labeling of 3D Volumes
Maximilian Söchting, Stefano Allegretti, Federico Bolelli, Costantino Grana

Auto-TLDR; Entropy Partitioning Decision Tree for Connected Components Labeling
Abstract Slides Poster Similar
MedZip: 3D Medical Images Lossless Compressor Using Recurrent Neural Network (LSTM)
Omniah Nagoor, Joss Whittle, Jingjing Deng, Benjamin Mora, Mark W. Jones

Auto-TLDR; Recurrent Neural Network for Lossless Medical Image Compression using Long Short-Term Memory
BG-Net: Boundary-Guided Network for Lung Segmentation on Clinical CT Images
Rui Xu, Yi Wang, Tiantian Liu, Xinchen Ye, Lin Lin, Yen-Wei Chen, Shoji Kido, Noriyuki Tomiyama

Auto-TLDR; Boundary-Guided Network for Lung Segmentation on CT Images
Abstract Slides Poster Similar
Revisiting Sequence-To-Sequence Video Object Segmentation with Multi-Task Loss and Skip-Memory
Fatemeh Azimi, Benjamin Bischke, Sebastian Palacio, Federico Raue, Jörn Hees, Andreas Dengel

Auto-TLDR; Sequence-to-Sequence Learning for Video Object Segmentation
Abstract Slides Poster Similar